RGraphSpace: A lightweight interface between igraph and ggplot2 graphics
Sysbiolab - Bioinformatics and Systems Biology Laboratory
2026-04-21
Source:vignettes/RGraphSpace.Rmd
RGraphSpace.RmdAbstract
RGraphSpace is an R package that integrates igraph objects with the ggplot2 ecosystem. It implements geometric prototypes designed for networks scaled to a standardized unit space. The package includes methods for side-by-side visualization of multiple graphs and for layering network elements onto spatial maps.
Package: RGraphSpace 1.1.3
Overview
RGraphSpace is an R package that generates ggplot2
graphics for igraph objects (Csardi and Nepusz
2006), scaling nodes and edges to a unit space. The package
implements new ggplot2 geometric prototypes (Wickham 2016), optimized for representing large
networks. This enables extensive customization of aesthetics and visual
style, including colors, shapes, and line types. Three specialized
geoms translate graph data into geometric layers (see using geoms). These geoms
use a dual-anchor normalization approach to align layers, which is
critical for analysis where network elements must be accurately
referenced to a spatial map. Section mapping graphs to images
illustrates how this alignment achieves pixel-level precision. In what
follows, this tutorial demonstrates how to use RGraphSpace for
side-by-side visualization of multiple graphs.
Quick start
This section will create a toy igraph object to demonstrate
the RGraphSpace workflow. The graph layout is configured
manually to ensure that users can easily view all the relevant arguments
needed to prepare the input data for the RGraphSpace package.
We will use the igraph’s make_star() function to create a
simple star-like graph and then the V() and
E() functions to set attributes for vertices and edges,
respectively. The RGraphSpace package will require that all
vertices have x, y, and name
attributes.
# Make a 'toy' igraph with 5 nodes and 4 edges;
# ..either a directed or undirected graph
gtoy1 <- make_star(5, mode="out")
# Check whether the graph is directed or not
is_directed(gtoy1)
## [1] TRUE
# Check graph size
vcount(gtoy1)
## [1] 5
ecount(gtoy1)
## [1] 4
# Assign 'x' and 'y' coordinates to each vertex;
# ..this can be an arbitrary unit in (-Inf, +Inf)
V(gtoy1)$x <- c(0, 2, -2, -4, -8)
V(gtoy1)$y <- c(0, 0, 2, -4, 0)
# Assign a name to each vertex
V(gtoy1)$name <- paste0("n", 1:5)
# Plot the 'gtoy1' using standard R graphics
plot(gtoy1)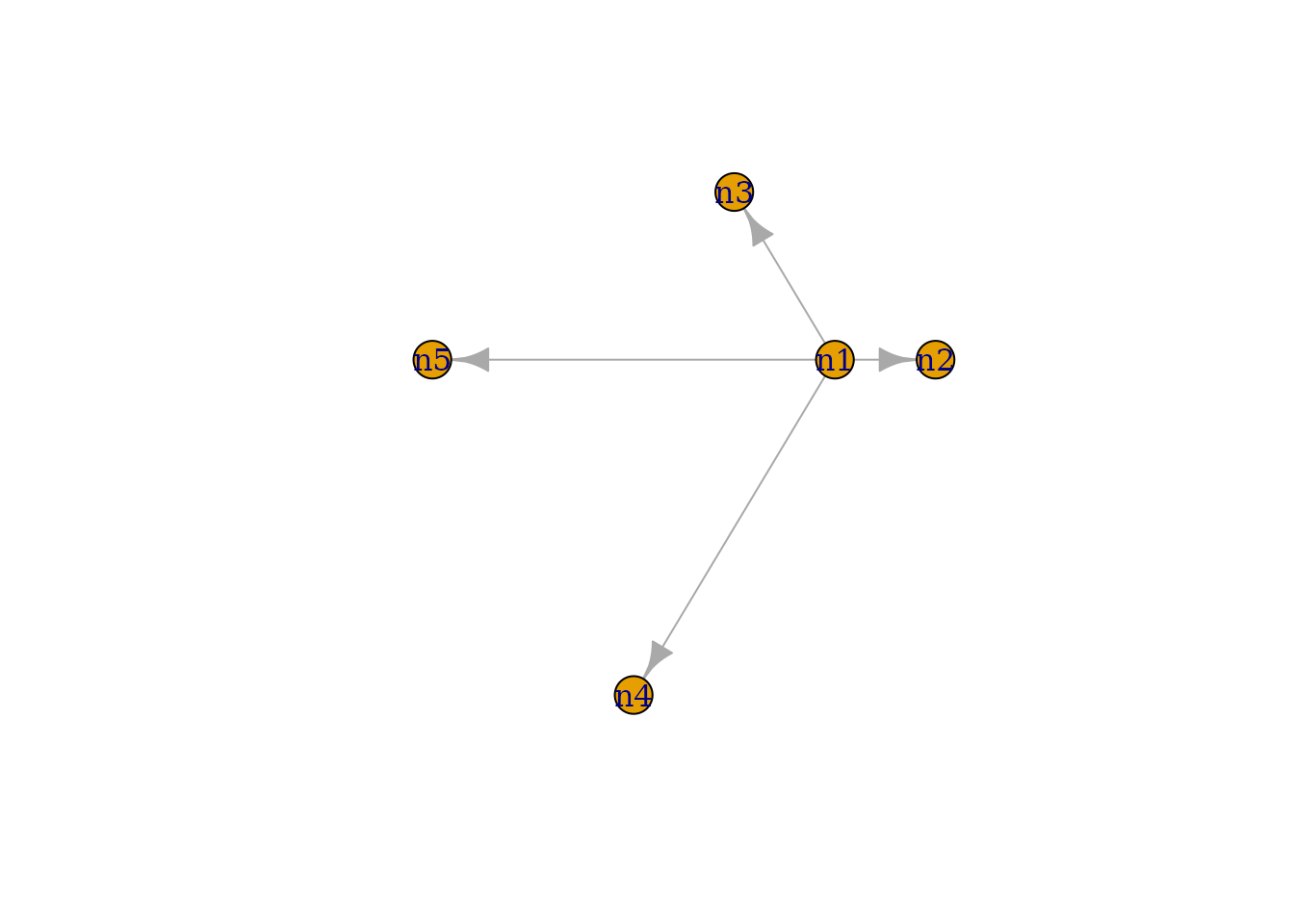
# Plot the 'gtoy1' using RGraphSpace
plotGraphSpace(gtoy1, add.labels = TRUE)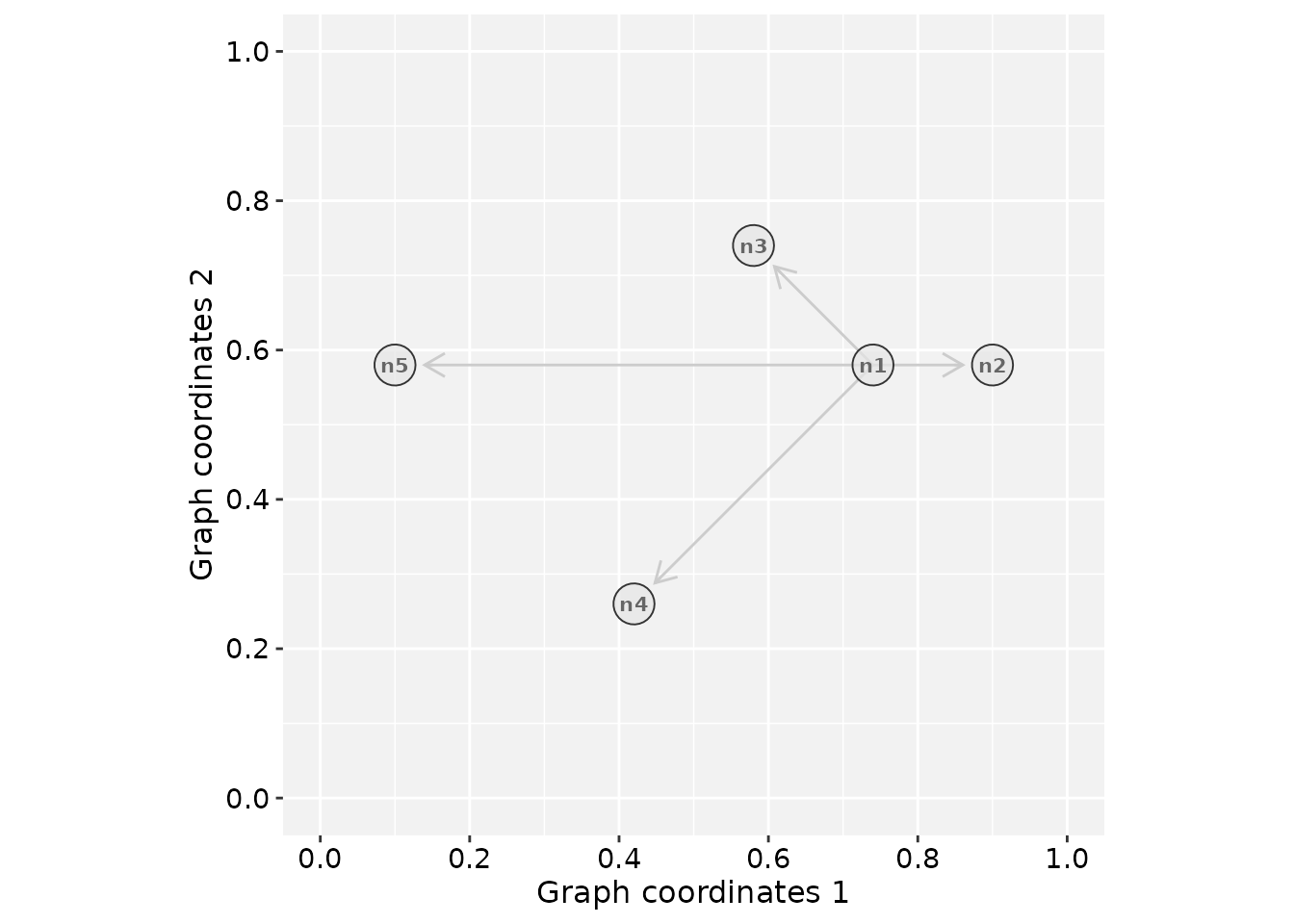
RGraphSpace attributes
Next, we will demonstrate all vertex and edge attributes that can be passed to RGraphSpace methods.
Vertex attributes
# Node size (numeric in [0, 100], as '%' of the plot space)
V(gtoy1)$nodeSize <- c(8, 5, 5, 10, 5)
# Node shape (integer code between 0 and 25; see 'help(points)')
V(gtoy1)$nodeShape <- c(21, 22, 23, 24, 25)
# Node color (Hexadecimal or color name)
V(gtoy1)$nodeColor <- c("red", "#00ad39", "grey80", "lightblue", "cyan")
# Node line width (as in 'lwd' standard graphics; see 'help(gpar)')
V(gtoy1)$nodeLineWidth <- 1
# Node line color (Hexadecimal or color name)
V(gtoy1)$nodeLineColor <- "grey20"
# Node labels ('NA' will omit labels)
V(gtoy1)$nodeLabel <- c("V1", "V2", "V3", "V4", NA)
# Node label size (in pts)
V(gtoy1)$nodeLabelSize <- 8
# Node label color (Hexadecimal or color name)
V(gtoy1)$nodeLabelColor <- "black"
# Node transparency (in [0,1])
V(gtoy1)$nodeAlpha <- 1Edge attributes
Given a list of edges, RGraphSpace represents only one edge for each pair of connected vertices. If there are multiple edges connecting the same vertex pairs, it will display the line attributes of the first edge in the list.
# Edge width (as in 'lwd' standard graphics; see 'help(gpar)')
E(gtoy1)$edgeLineWidth <- 0.8
# Edge color (Hexadecimal or color name)
E(gtoy1)$edgeLineColor <- c("red","green","blue","black")
# Edge type (as in 'lty' standard graphics; see 'help(gpar)')
E(gtoy1)$edgeLineType <- c("solid", "11", "dashed", "2124")
# Edge transparency (in [0,1])
E(gtoy1)$edgeAlpha <- 1Arrowhead attributes
Arrowhead in directed graphs: By default, an arrow will be drawn for
each edge according to its left-to-right orientation in the edge list
(e.g. A -> B).
# Arrowhead types in directed graphs (integer code or character)
## 0 = "---", 1 = "-->", -1 = "--|"
E(gtoy1)$arrowType <- 1Arrowhead in undirected graphs: By default, no arrow will be drawn in undirected graphs.
# Arrowhead types in undirected graphs (integer or character code)
## 0 = "---"
## 1 = "-->", 2 = "<--", 3 = "<->", 4 = "|->",
## -1 = "--|", -2 = "|--", -3 = "|-|", -4 = "<-|",
E(gtoy1)$arrowType <- 1
# Note: in undirected graphs, this attribute overrides the
# edge's orientation in the edge list… and now plot the updated igraph object with RGraphSpace:
# Plot the updated 'gtoy1' using RGraphSpace
plotGraphSpace(gtoy1, add.labels = TRUE)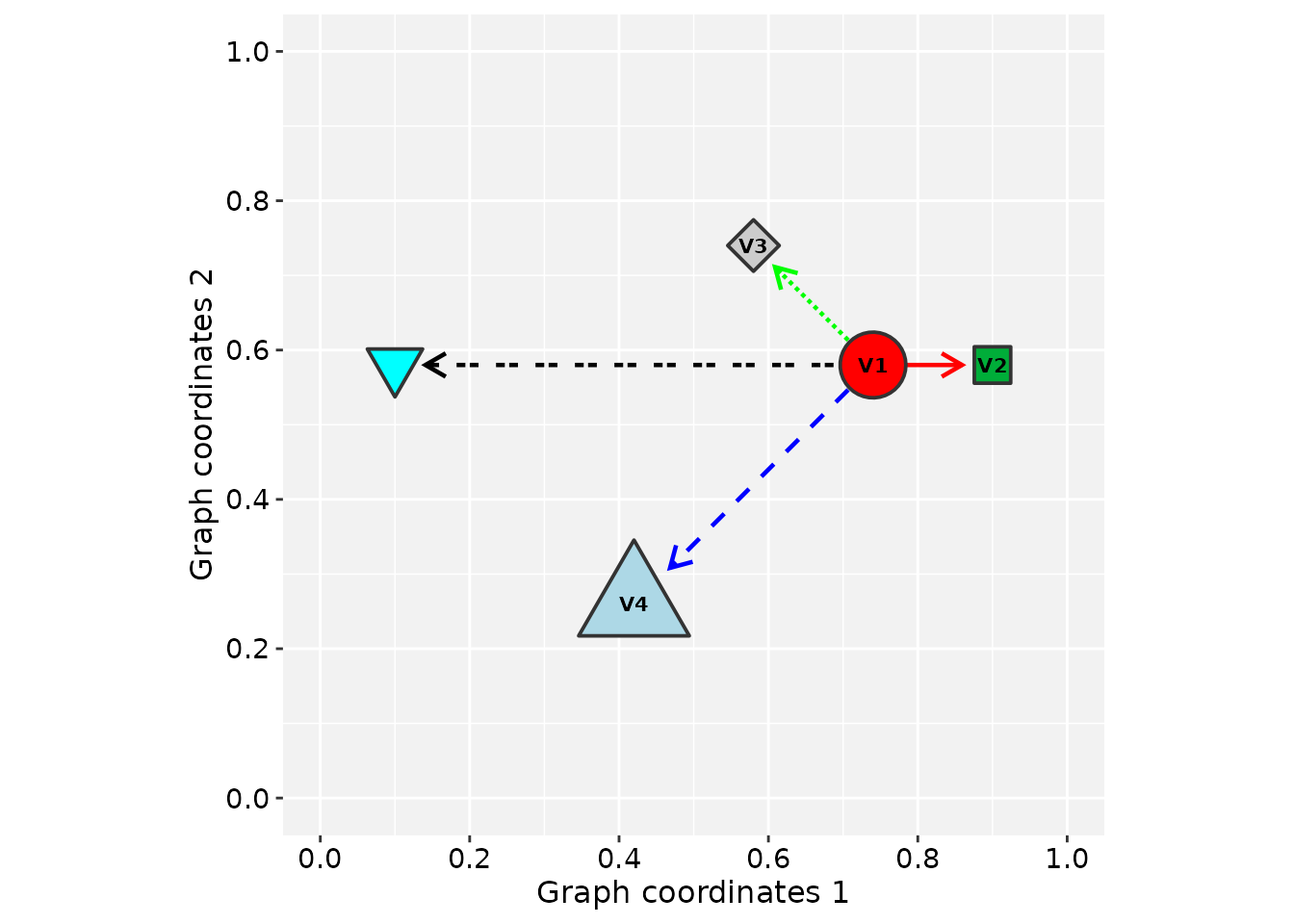
Using ggplot2 geoms
Visual integration and aesthetics mapping
This section illustrates how RGraphSpace integrates with the
ggplot2 using geoms building blocks. Graph
attributes stored within the GraphSpace object can be
handled in two ways:
Identity Mapping: Graph attributes are interpreted as “identity values” (such as
nodeColor,nodeSize, ornodeShape) and are displayed exactly as they are, without further scaling or mapping.Dynamic Aesthetic Mapping: Graph attributes are mapped to aesthetics (such as
colour,size, andshape) and rendered through standard ggplot2 scales, which automatically generate synchronized legends.
The GraphSpace geoms
To facilitate this integration, RGraphSpace implements three
specialized geoms designed to handle graph data types
within a ggplot2 workflow:
-
geom_graphspace(): A high-level convenience layer that processes both nodes and edges in a single call. -
geom_nodespace(): Dedicated to rendering nodes. InheritsGeomPointaesthetic mappings, optimized to inform the edge layer on node states. It is used with theinject_nodespace()function to adjust edge offsets. -
geom_edgespace(): Handles the relational data between nodes. InheritsGeomSegmentaesthetic mappings; unlike standard segment geoms, it is “node-aware” and dynamically adjusts geometries based on connected nodes.
In the following example, we create a small modular graph containing variables of different types in order to demonstrate these geoms.
# Make a toy modular graph
library("igraph")
gtoy3 <- sample_islands(
islands.n = 3, # number of modules
islands.size = 30, # nodes per module
islands.pin = 0.25, # probability of edges within modules
n.inter = 2) # edges between modules
# Assign module membership to nodes
V(gtoy3)$module <- rep(1:3, each = 30)
# Assign colors to nodes
V(gtoy3)$nodeColor <- rainbow(3)[V(gtoy3)$module]
# Assign a categorical variable to nodes
V(gtoy3)$node_group <- c("A", "B", "C")[V(gtoy3)$module]
# Assign numeric variables to nodes and edges
V(gtoy3)$node_var <- runif(vcount(gtoy3))
E(gtoy3)$edge_var <- runif(ecount(gtoy3))
# Create a GraphSpace from the toy igraph
gs <- GraphSpace(gtoy3)Plotting identity values
In this example, nodeColor already contains the final
colour values stored in the GraphSpace object. The colours
will be displayed as-is by the geom_graphspace() function.
This approach is particularly useful when nodes have been pre-processed
with specific color schemes and you want the visual output to reflect
the original data directly, without further mapping.
ggplot() +
geom_graphspace(colour = "grey", data = gs) +
theme(aspect.ratio = 1)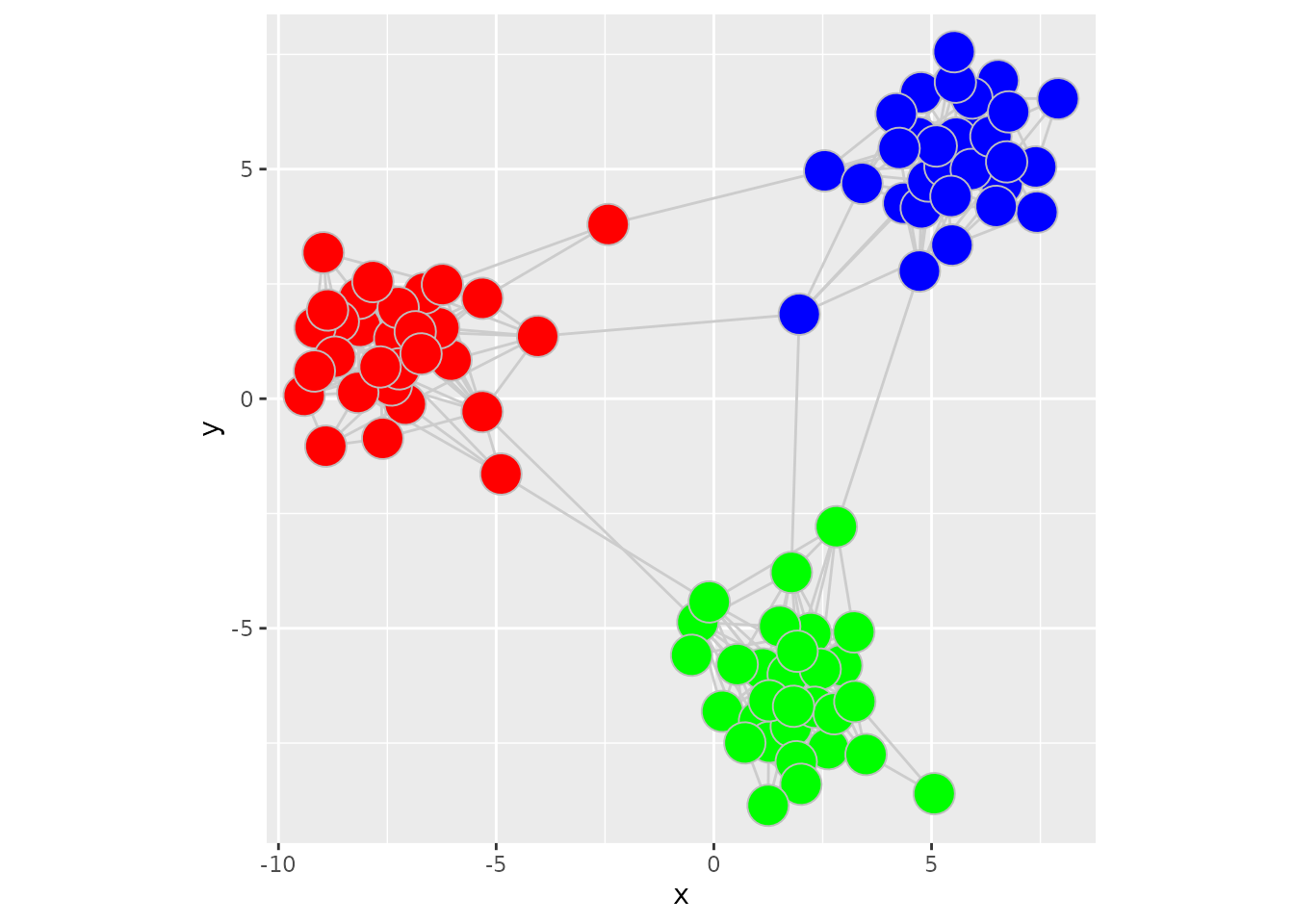
The trade-off on this approach is that, on one hand, all attributes
remain scaled with the graph space, but no legend is accessible. This is
because identity scales bypass the scaling and guide-building process of
ggplot2. If a legend is required to explain the meaning
of these colors, the attribute should be mapped as a variable (e.g.,
aes(fill = attribute)) using standard discrete or
continuous scales.
Mapping categorical variables
In this example, the node categorical variable
node_group is mapped to the fill
aesthetic.
ggplot() +
geom_graphspace(aes(fill = node_group),
colour = "grey", data = gs) +
scale_fill_viridis_d(option = "viridis") +
theme_gspace_coords()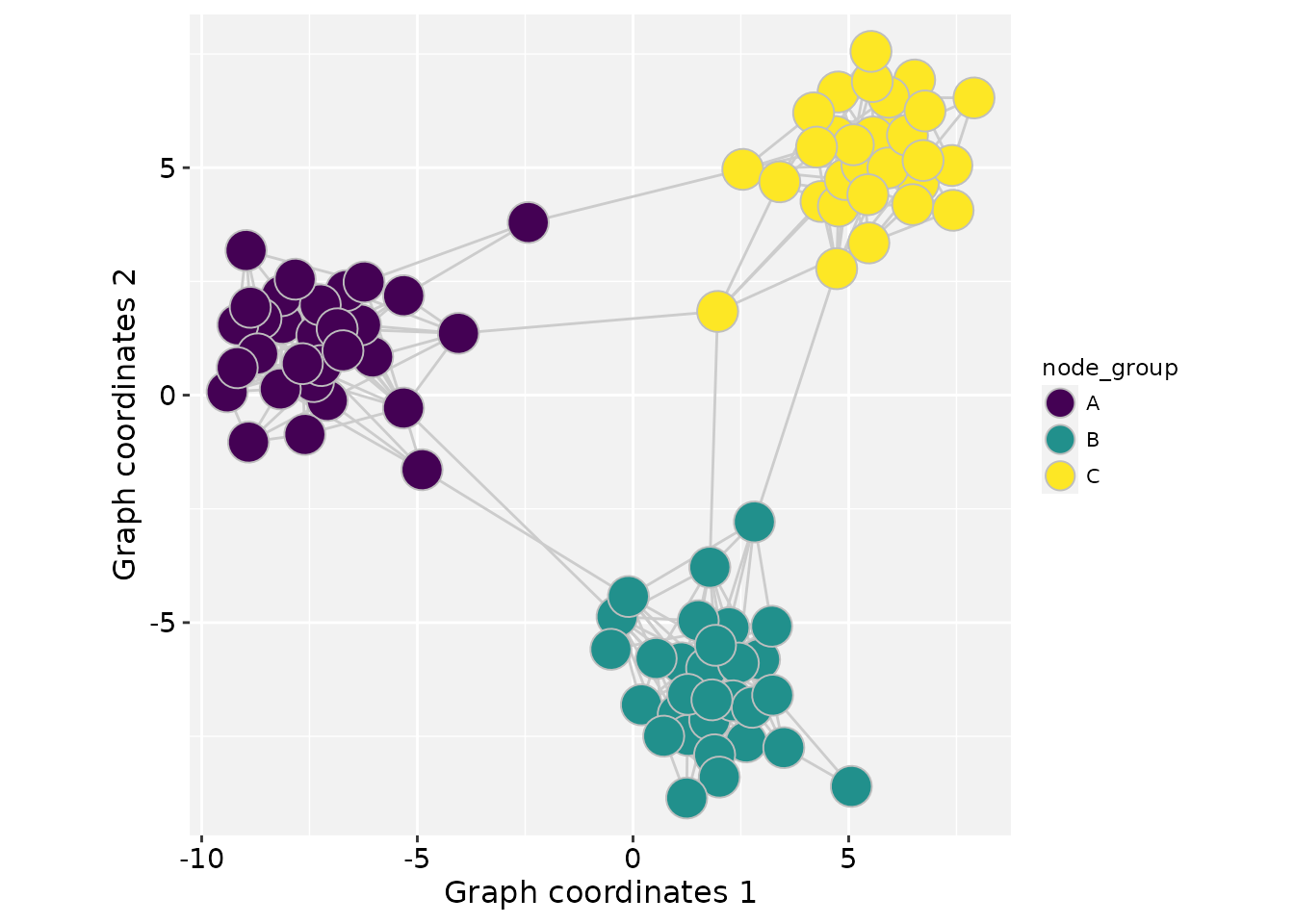
Mapping numeric variables
In this example, node and edge numeric variables are mapped to
fill and colour aesthetics, respectively.
# Map aesthetics to numeric variables
ggplot() +
geom_edgespace(aes(colour = edge_var), data = gs) +
geom_nodespace(aes(fill = node_var),
colour = "grey", data = gs) +
scale_colour_continuous(palette = c("cyan","blue")) +
scale_fill_continuous(palette = c("white","purple")) +
theme_gspace_coords()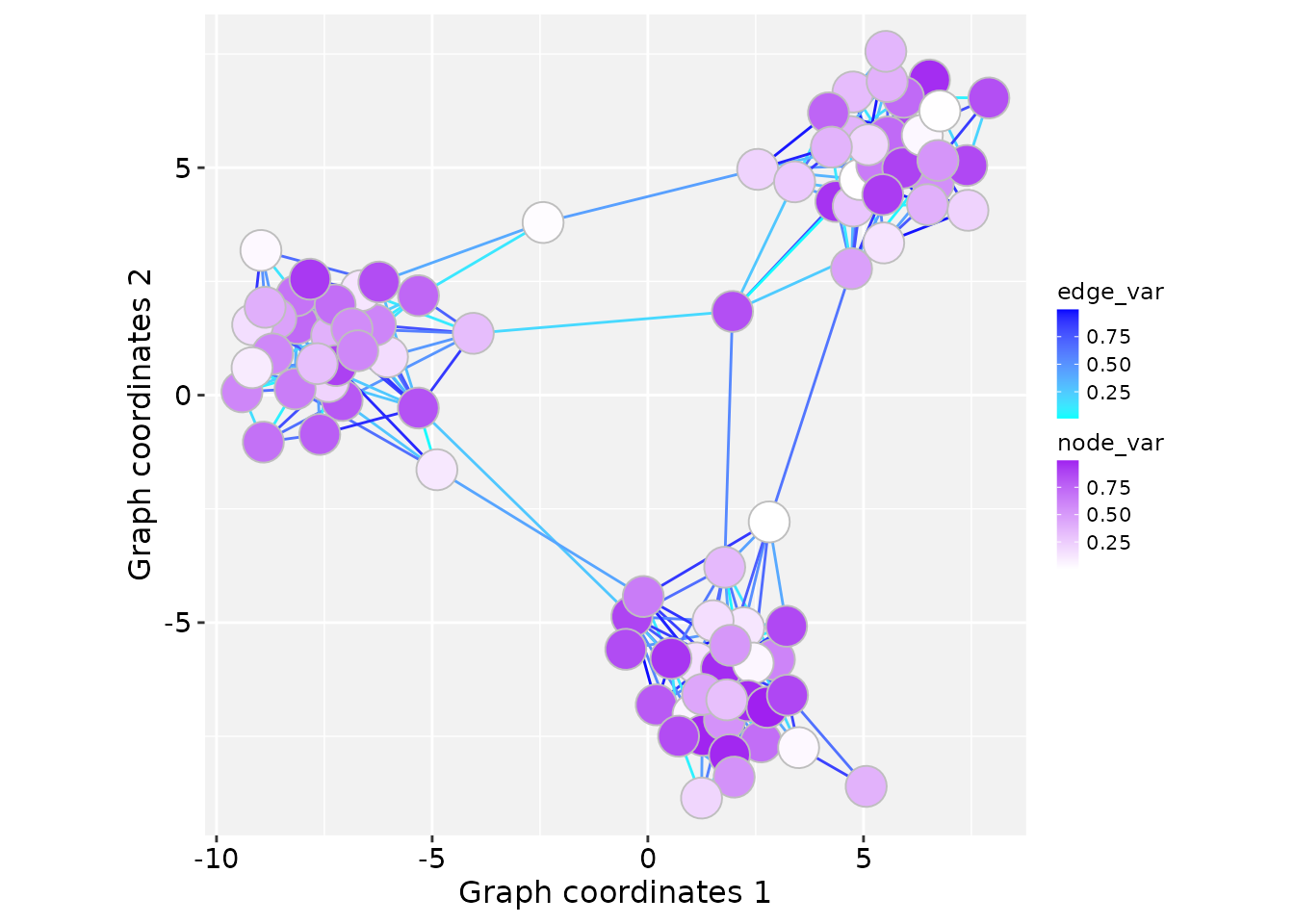
Using separate colour scales
When multiple geoms use the same aesthetic (for example
colour) but require mapping to different variables with
independent scales, the ggnewscale package can be used to
introduce a new scale (Campitelli
2025).
if (!require("ggnewscale", quietly = TRUE)) {
install.packages("ggnewscale")
}
library("ggnewscale")
ggplot() +
geom_edgespace(aes(colour = edge_var), data = gs) +
scale_colour_continuous(palette = c("cyan","blue")) +
ggnewscale::new_scale_colour() +
geom_nodespace(aes(colour = node_var),
data = gs, stroke = 2, fill = NA) +
scale_colour_continuous(palette = c("white","purple")) +
theme_gspace_coords()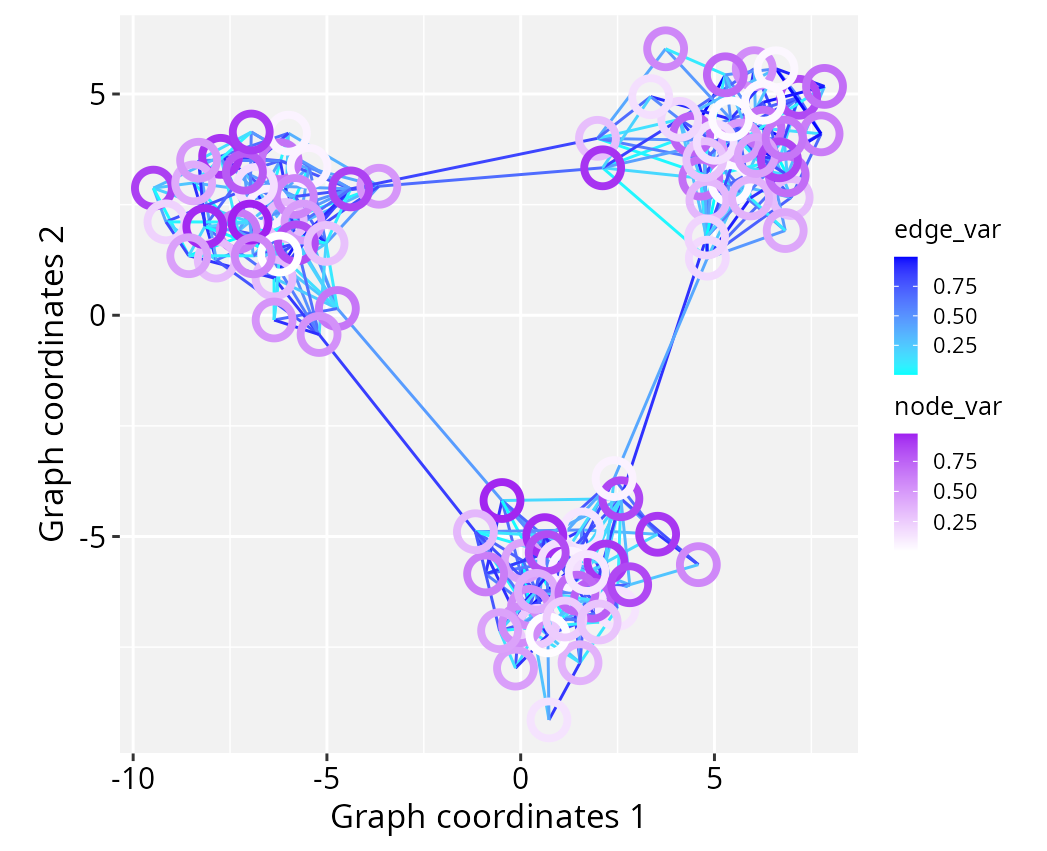
Mapping graphs to images
Images can be used as spatial references for graph layouts in RGraphSpace. When a raster image is provided, its pixel grid defines the coordinate system where nodes are positioned, supporting the construction of graphs from image features.
Next, we use the volcano topographic matrix as an
example. Features extracted from this matrix are mapped to graph nodes
and visualized over a raster background.
# Extract pixel coordinates for a specific intensity quantile.
coords <- which(volcano == quantile(volcano, 0.85), arr.ind = TRUE)
# Mark target pixels with '0'; it will appear as black in the background.
# This creates a visual anchor to verify the alignment precision.
volcano2 <- volcano
volcano2[coords] <- 0
# Create an igraph object from the pixel coordinates;
# note that at this stage, 'y' represents matrix row indices.
gtoy2 <- igraph::make_empty_graph(n = nrow(coords))
igraph::V(gtoy2)$y <- coords[,1]
igraph::V(gtoy2)$x <- coords[,2]
# Highlight the bottom-row vertex (max 'y' index) to demonstrate alignment;
# since matrix indexing is top-down, this accounts for the default flip
# between matrix and plot coordinate systems.
igraph::V(gtoy2)$nodeColor <- NA
bottom_row <- which.max(igraph::V(gtoy2)$y)
igraph::V(gtoy2)$nodeColor[bottom_row] <- "red"
# Initialize a GraphSpace object
gs <- GraphSpace(gtoy2)
# Map graph coordinates to the image space.
# Vertical flipping (flip.v = TRUE) is required to align coordinates.
gs <- normalizeGraphSpace(gs, image = as_colorraster(volcano2),
flip.v = TRUE)
# Render the graph with the raster as background
plotGraphSpace(gs, add.image = TRUE)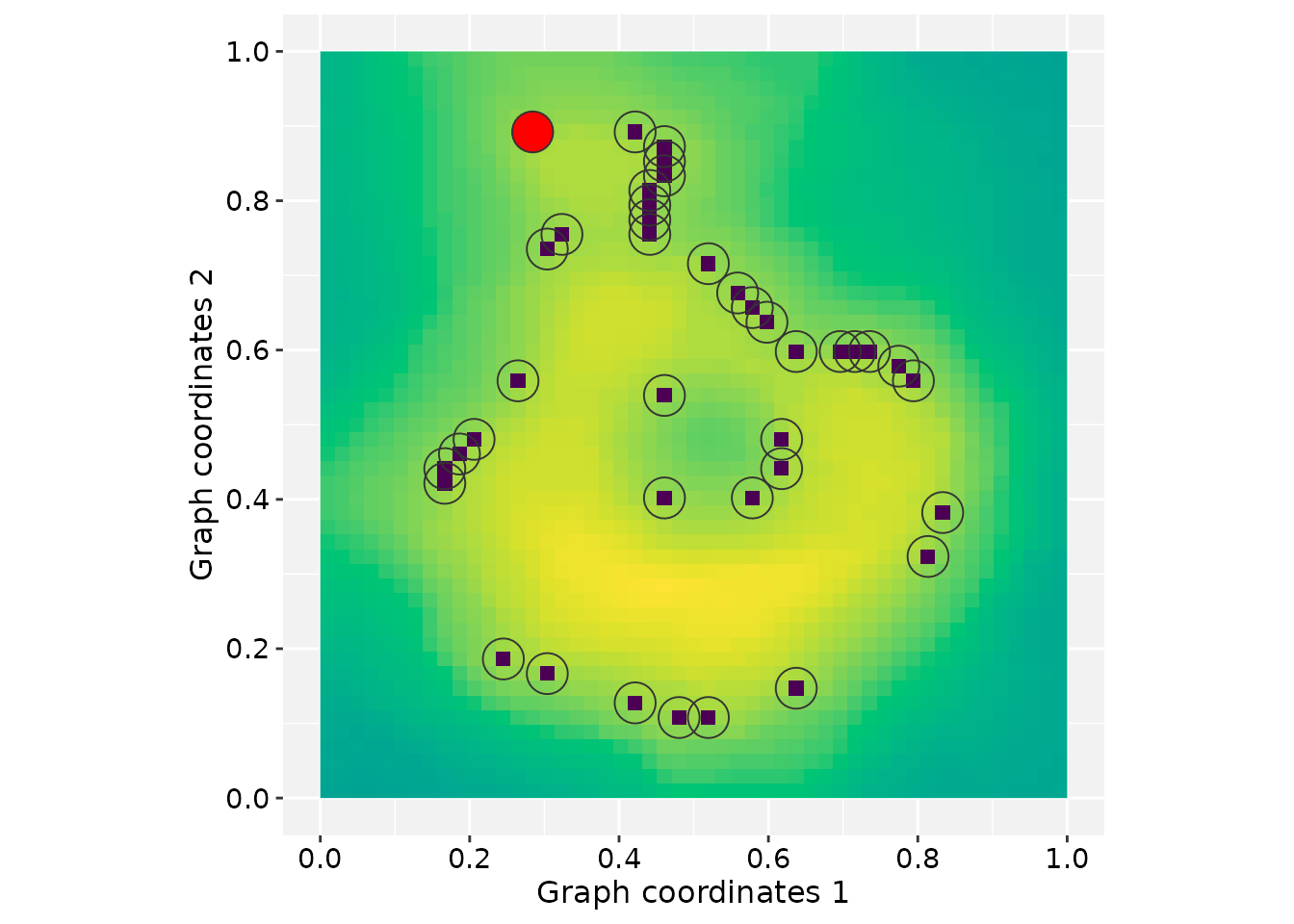
Interoperability with other packages
Geospatial data
The following example demonstrates interoperability between RGraphSpace and sf, a well-established infrastructure package for spatial data analysis (Pebesma and Bivand 2023). We will use a spatial network of cities to show how RGraphSpace geoms can be plugged into sf workflows.
if(!require("sf", quietly = TRUE)){
install.packages("sf")
}
if(!require("rnaturalearth", quietly = TRUE)){
install.packages("rnaturalearth")
}
if(!require("maps", quietly = TRUE)){
install.packages("maps")
}
if(!require("geometry", quietly = TRUE)){
install.packages("geometry")
}
library("RGraphSpace")
library("igraph")
library("sf")
library("maps")
library("geometry")
library("rnaturalearth")
# Load and project map
map_sf <- ne_countries(country = "Brazil", returnclass = "sf")
map_proj <- st_transform(map_sf)
# Filter major cities by regional capitals
data(world.cities, package = "maps")
r_capitals <- c(
"Aracaju", "Belem", "Belo Horizonte", "Boa Vista", "Brasilia",
"Campo Grande", "Cuiaba", "Curitiba", "Florianopolis", "Fortaleza",
"Goiania", "Joao Pessoa", "Macapa", "Maceio", "Manaus", "Natal",
"Palmas", "Porto Alegre", "Porto Velho", "Recife", "Rio Branco",
"Rio de Janeiro", "Salvador", "Sao Luis", "Sao Paulo", "Teresina",
"Vitoria"
)
cities <- subset(world.cities, country.etc == "Brazil" &
name %in% r_capitals & pop > 1000000)
# Create Delaunay triangulation edges
# Note: the edges hold no particular meaning beyond
# demonstrating integration between coordinate systems
tri <- delaunayn(cities[,c("lat","long")])
edges <- unique(rbind(tri[,c(1,2)], tri[,c(2,3)], tri[,c(1,3)] ))
# Build igraph with coordinates
gtoy1 <- igraph::graph_from_edgelist(edges, directed = FALSE)
igraph::V(gtoy1)$x <- cities$long
igraph::V(gtoy1)$y <- cities$lat
igraph::V(gtoy1)$Cities <- cities$name
igraph::V(gtoy1)$`Population (M)` <- cities$pop/1000000
igraph::E(gtoy1)$arrowType <- 3
# Make a GraphSpace
gs1 <- GraphSpace(gtoy1)
# Plot
ggplot() +
geom_sf(data = map_proj, fill = "grey95", color = "grey60") +
geom_edgespace(color = "grey40", arrow_size = 0.5,
arrow_offset = 0.01, data = gs1) +
geom_nodespace(aes(fill = Cities, size = `Population (M)`),
data = gs1) +
scale_size(range = c(3,9)) +
scale_fill_discrete() +
inject_nodespace() +
theme_gspace_legend(key_fill = TRUE)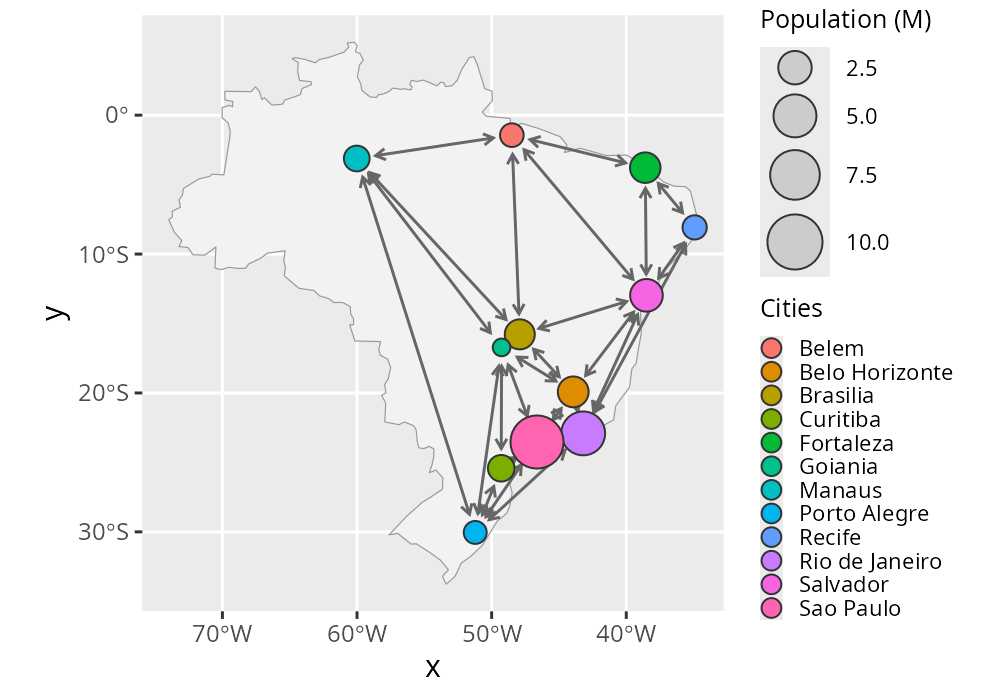
Interactive visualization
The following example demonstrates interoperability between RGraphSpace and RedeR, an R/Bioconductor package for interactive network visualization and manipulation.
# Load RedeR, a graph package for interactive visualization
## Note: this example requires Bioc >= 3.19
if(!require("BiocManager", quietly = TRUE)){
install.packages("BiocManager")
#BiocManager::install(version = "3.19")
}
if(!require("RedeR", quietly = TRUE)){
BiocManager::install("RedeR")
}
# Launch the RedeR application
library("RedeR")
startRedeR()
resetRedeR()
data(gtoy1, package = "RGraphSpace")
# Send 'gtoy1' to the RedeR interface
addGraphToRedeR(gtoy1, unit="npc")
relaxRedeR()
# Fetch 'gtoy1' with a fresh layout
gtoy2 <- getGraphFromRedeR(unit="npc")
# Check the round trip...
plotGraphSpace(gtoy2, add.labels = TRUE)
## Note that for the round trip, shapes and line types are
## partially compatible between ggplot2 and RedeR.
# ...alternatively, just update the graph layout
gtoy2 <- updateLayoutFromRedeR(g=gtoy1)
# ...check the updated layout
plotGraphSpace(gtoy2, add.labels = TRUE)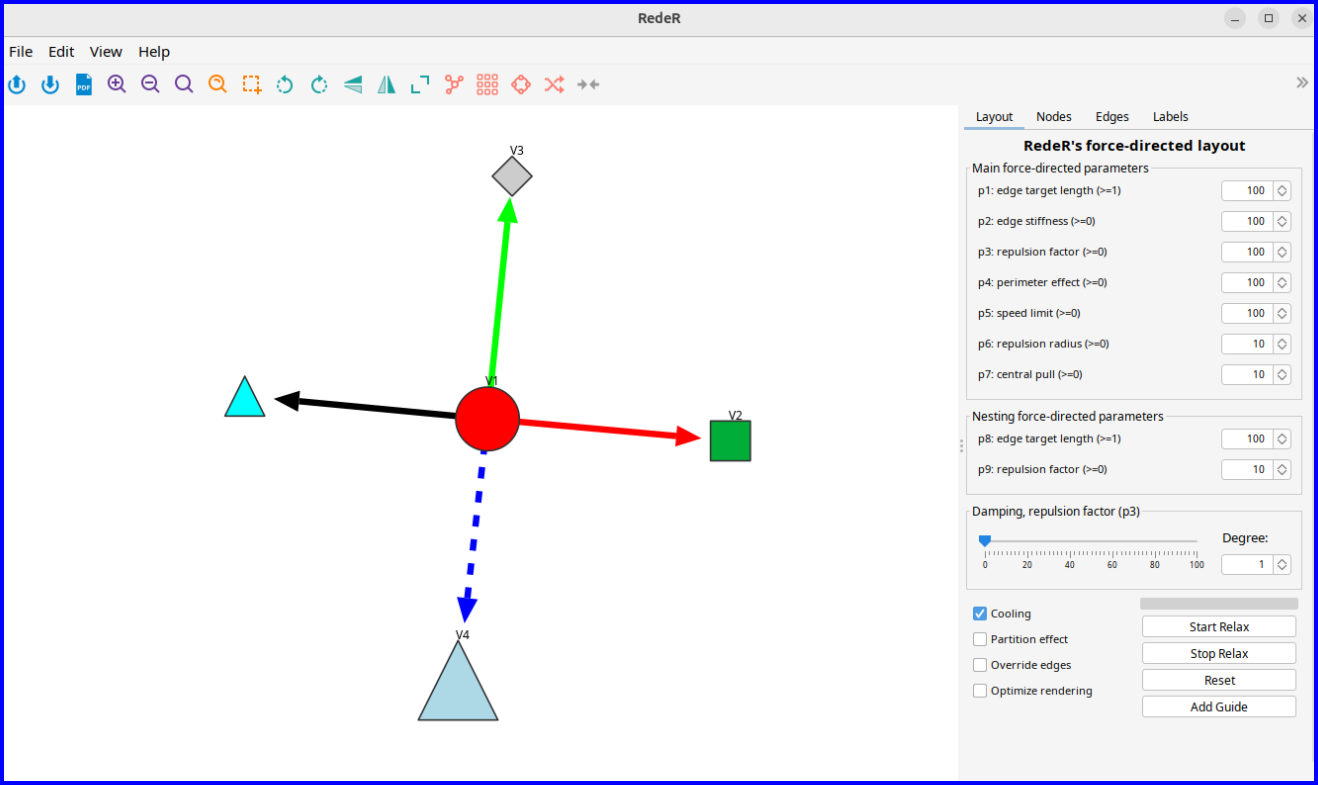
Citation
If you use RGraphSpace, please cite:
Sysbiolab Team. “RGraphSpace: A lightweight interface between igraph and ggplot2 graphics.” R package, 2023. Doi: 10.32614/CRAN.package.RGraphSpace
Castro MA, Wang X, Fletcher MN, Meyer KB, Markowetz F (2012). “RedeR: R/Bioconductor package for representing modular structures, nested networks and multiple levels of hierarchical associations.” Genome Biology, 13(4), R29. Doi: 10.1186/gb-2012-13-4-r29
Session information
#> R version 4.5.3 (2026-03-11)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] RGraphSpace_1.1.3 ggplot2_4.0.2 igraph_2.2.3
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.5.3 ggbeeswarm_0.7.3
#> [5] jquerylib_0.1.4 systemfonts_1.3.2 scales_1.4.0 textshaping_1.0.5
#> [9] yaml_2.3.12 fastmap_1.2.0 R6_2.6.1 labeling_0.4.3
#> [13] knitr_1.51 desc_1.4.3 bslib_0.10.0 RColorBrewer_1.1-3
#> [17] rlang_1.2.0 cachem_1.1.0 xfun_0.57 fs_2.1.0
#> [21] sass_0.4.10 S7_0.2.1-1 viridisLite_0.4.3 cli_3.6.6
#> [25] pkgdown_2.2.0 withr_3.0.2 magrittr_2.0.5 digest_0.6.39
#> [29] grid_4.5.3 beeswarm_0.4.0 lifecycle_1.0.5 ggrastr_1.0.2
#> [33] vipor_0.4.7 vctrs_0.7.3 evaluate_1.0.5 glue_1.8.1
#> [37] farver_2.1.2 ragg_1.5.2 rmarkdown_2.31 tools_4.5.3
#> [41] pkgconfig_2.0.3 htmltools_0.5.9